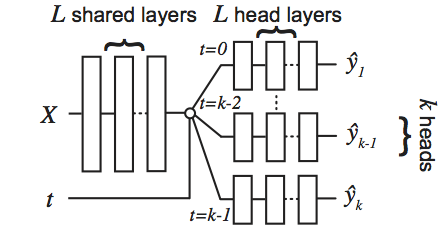
Perfect Match: A Simple Method for Learning Representations For Counterfactual Inference With Neural Networks
Perfect Match (PM) is a method for learning to estimate individual treatment effect (ITE) using neural networks. PM is easy to implement, compatible with any architecture, does not add computational complexity or hyperparameters, and extends to any number of treatments. This repository contains the source code used to evaluate PM and most of the existing state-of-the-art methods at the time of publication of our manuscript. PM and the presented experiments are described in detail in our paper. Since we performed one of the most comprehensive evaluations to date with four different datasets with varying characteristics, this repository may serve as a benchmark suite for developing your own methods for estimating causal effects using machine learning methods. In particular, the source code is designed to be easily extensible with (1) new methods and (2) new benchmark datasets.
Author(s): Patrick Schwab, ETH Zurich patrick.schwab@hest.ethz.ch, Lorenz Linhardt, ETH Zurich llorenz@student.ethz.ch and Walter Karlen, ETH Zurich walter.karlen@hest.ethz.ch
License: MIT, see LICENSE.txt
If you reference or use our methodology, code or results in your work, please consider citing:
@article{schwab2018perfect,
title={{Perfect Match: A Simple Method for Learning Representations For Counterfactual Inference With Neural Networks}},
author={Schwab, Patrick and Linhardt, Lorenz and Karlen, Walter},
journal={arXiv preprint arXiv:1810.00656},
year={2018}
}
- Runnable scripts are in the
perfect_match/apps/subdirectory.perfect_match/apps/main.pyis the main runnable script for running experiments.- The available command line parameters for runnable scripts are described in
perfect_match/apps/parameters.py
- You can add new baseline methods to the evaluation by subclassing
perfect_match/models/baselines/baseline.py- See e.g.
perfect_match/models/baselines/neural_network.pyfor an example of how to implement your own baseline methods. - You can register new methods for use from the command line by adding a new entry to the
get_method_name_mapmethod inperfect_match/apps/main.py
- See e.g.
- You can add new benchmarks by implementing the benchmark interface, see e.g.
perfect_match/models/benchmarksfor examples of how to add your own benchmark to the benchmark suite.- You can register new benchmarks for use from the command line by adding a new entry to the
get_benchmark_name_mapmethod inperfect_match/apps/evaluate.py
- You can register new benchmarks for use from the command line by adding a new entry to the
-
This project was designed for use with Python 2.7. We can not guarantee and have not tested compability with Python 3.
-
To run the IHDP benchmark, you need to download the raw IHDP data folds as used by Johanson et al. available at this link.
- After downloading IHDP-1000.tar.gz, you must extract the files into the
./perfect_match/data_access/ihdpdirectory relative to this file. - e.g. using:
tar xvzf IHDP-1000.tar.gz -C ./perfect_match/data_access/ihdp - Ensure that the files
ihdp_npci_1-1000.train.npzandihdp_npci_1-1000.test.npzare present in the./perfect_match/data_access/ihdpdirectory.
- After downloading IHDP-1000.tar.gz, you must extract the files into the
-
To run the TCGA and News benchmarks, you need to download the SQLite databases containing the raw data samples for these benchmarks (
news.dbandtcga.db).- You can download the raw data under these links: tcga.db and news.db.
- Note that you need around 10GB of free disk space to store the databases.
- Save the database files to the
./datadirectory relative to this file in order to be compatible with the step-by-step guides below or adjust the commands accordingly.
-
To run BART, Causal Forests and to reproduce the figures you need to have R installed. See https://www.r-project.org/ for installation instructions.
- To run BART, you need to have the R-packages
rJavaandbartMachineinstalled. See https://github.com/kapelner/bartMachine for installation instructions. Note thatrJavarequires a working Java installation as well. - To run Causal Forests, you need to have the R-package
grfinstalled. See https://github.com/grf-labs/grf for installation instructions. - To reproduce the paper's figures, you need to have the R-package
latex2expinstalled. See https://cran.r-project.org/web/packages/latex2exp/vignettes/using-latex2exp.html for installation instructions.
- To run BART, you need to have the R-packages
-
For the python dependencies, see
setup.py. You can usepip install .to install the perfect_match package and the python dependencies. Note the installation ofrpy2will fail if you do not have a working R installation on your system (see above).
- Make sure you have all the requirements listed above.
- You can use the script
perfect_match/apps/run_all_experiments.pyto obtain the exact parameters used withmain.pyto reproduce the experimental results in our paper.- The
perfect_match/apps/run_all_experiments.pyscript prints the command line parameters that have to be run, one command per line. - Note that we ran several thousand experiments which can take a while if evaluated sequentially. We therefore suggest to run the commands in parallel using, e.g., a compute cluster.
- The original experiments reported in our paper were run on Intel CPUs. We found that running the experiments on GPUs can produce ever so slightly different results for the same experiments.
- The
- Once you have completed the experiments, you can calculate the summary statistics (mean +- standard deviation) over all the repeated runs using the
./run_results.shscript. The results are reported in LaTeX syntax in the order reported in the results tables, i.e. {12.2}$\pm$ 0.1 & {14.3}$\pm$ 0.2 where 12.2, and 14.3 are the means of PEHE, and ATE (for News and IHDP) and the R_Pol and ATT (for Jobs) and 0.1 and 0.2 are the standard deviations, respectively.- See below for a step-by-step guide for each reported result.
- If the
./run_results.shscript produces errors, one or multiple of your runs may have failed to complete successfully. You can check the run's run.txt file to see whether there have been any errors.
- You can also reproduce the figures in our manuscript by running the R-scripts in
perfect_match/visualisation/
- Navigate to the directory containing this file.
- Create a folder to hold the experimental results
mkdir -p results. - Run
python ./perfect_match/apps/run_all_experiments.py ./perfect_match/apps ihdp ./data ./results- The script will print all the command line configurations (13000 in total) you need to run to obtain the experimental results to reproduce the IHDP results.
- Run the command line configurations from the previous step in a compute environment of your choice.
- After the experiments have concluded, use
./run_results.shto calculate the summary statistics mean +- standard deviation over all repeated runs in LaTeX syntax.- Use
./run_results.sh ./results/pm_ihdp2a0k_{METHOD_NAME}_mse_1, where {METHOD_NAME} should be replaced with the shorthand code of the method for which you wish to read out the result metrics. You can find the mapping of available shorthand codes to methods in theperfect_match/apps/main.pyfile. - Example 1:
./run_results.sh ./results/pm_ihdp2a0k_pbm_mse_1 ihdp, whereihdpindicates that you want results for the IHDP dataset, to get the results for "PM" on IHDP. - Example 2:
./run_results.sh ./results/pm_ihdp2a0k_pbm_mahal_mse_1 ihdpto get the results for "+ on X" on IHDP. - Example 3:
./run_results.sh ./results/pm_ihdp2a0k_pbm_no_tarnet_mse_1 ihdpto get the results for "+ MLP" on IHDP.
- Use
- Navigate to the directory containing this file.
- Create a folder to hold the experimental results
mkdir -p results. - Run
python ./perfect_match/apps/run_all_experiments.py ./perfect_match/apps jobs ./data ./results- The script will print all the command line configurations (40 in total) you need to run to obtain the experimental results to reproduce the Jobs results.
- Note that we only evaluate PM, + on X, + MLP, PSM on Jobs. All other results are taken from the respective original authors' manuscripts.
- Run the command line configurations from the previous step in a compute environment of your choice.
- After the experiments have concluded, use
./run_results.shto calculate the summary statistics mean +- standard deviation over all repeated runs in LaTeX syntax.- Example 1:
./run_results.sh ./results/pm_jobs2a0k_pbm_mse_1 jobs, wherejobsindicates that you want results for the jobs dataset, to get the results for "PM" on Jobs. - Example 2:
./run_results.sh ./results/pm_jobs2a0k_pbm_mahal_mse_1 jobsto get the results for "+ on X" on Jobs. - Example 3:
./run_results.sh ./results/pm_jobs2a0k_pbm_no_tarnet_mse_1 jobsto get the results for "+ MLP" on Jobs. - Example 4:
./run_results.sh ./results/pm_jobs2a0k_psm_mse_1 jobsto get the results for "PSM" on Jobs.
- Example 1:
- Navigate to the directory containing this file.
- Create a folder to hold the experimental results
mkdir -p results. - Run
python ./perfect_match/apps/run_all_experiments.py ./perfect_match/apps news ./data ./results- The script will print all the command line configurations (2400 in total) you need to run to obtain the experimental results to reproduce the News results.
- Run the command line configurations from the previous step in a compute environment of your choice.
- After the experiments have concluded, use
./run_results.shto calculate the summary statistics mean +- standard deviation over all repeated runs in LaTeX syntax.- Example 1:
./run_results.sh ./results/pm_news2a10k_pbm_mse_1 news-2, wherenews-2indicates that you want results for the News-2 dataset, to get the results for "PM" on News-2. Note that the folder path must match exactly with the type of dataset requested (news-2 <> news-2), otherwise the shown summary statistics will not be the right metrics. - Example 2:
./run_results.sh ./results/pm_news4a10k_pbm_mse_1 news-4to get the results for "PM" on News-4. - Example 3:
./run_results.sh ./results/pm_news8a10k_pbm_mse_1 news-8to get the results for "PM" on News-8. - Example 4:
./run_results.sh ./results/pm_news16a7k_pbm_mse_1 news-16to get the results for "PM" on News-16. - Repeat for all evaluated method / benchmark combinations.
- Example 1:
- Go through the IHDP step-by-step above.
- Ensure that you have run the
./run_results.shscript at least once on./results/pm_ihdp2a0k_pbm_mse_1and that the summary.txt file was created in./results/pm_ihdp2a0k_pbm_mse_1. - Navigate to the
./results/pm_ihdp2a0k_pbm_mse_1directory. - Run the following scripts to obtain mse.txt, pehe.txt and nn_pehe.txt for use with the
perfect_match/visualisation/cor_plots.Rscript:cat summary.txt | grep "val_f MSE" | awk '{print $7}' > mse.txtcat summary.txt | grep "val_pehe" | awk '{print $7}' > pehe.txtcat summary.txt | grep "val_pehe" | awk '{print $10}' > nn_pehe.txt
- You can use the
perfect_match/visualisation/corr_plot.Rfile to reproduce the correlation plots in Figure 1.
- Navigate to the directory containing this file.
- Create a folder to hold the experimental results
mkdir -p results. - Run
python ./perfect_match/apps/run_all_experiments.py ./perfect_match/apps news_matching_percentage ./data ./results- The script will print all the command line configurations (450 in total) you need to run to obtain the experimental results to reproduce the News results.
- Run the command line configurations from the previous step in a compute environment of your choice.
- After the experiments have concluded, use
./run_results.shto calculate the summary statistics mean +- standard deviation over all repeated runs in LaTeX syntax.- Example 1:
./run_results.sh ./results/pm_news8a10k0.10p_pbm_mse_1 news-8, wherenews-8indicates that you want results for the News-8 dataset and0.10pindicates 10% matched samples per batch, to get the results for "PM" on News-8 with 10% matched samples per batch. - Example 2:
./run_results.sh ./results/pm_news8a10k0.20p_pbm_mse_1 news-8to get the results for "PM" on News-8 with 20% matched samples per batch. - Repeat for all evaluated percentages of matched samples.
- Example 1:
- Your results should match those found in the
perfect_match/visualisation/percentage_plot.Rfile.
- Navigate to the directory containing this file.
- Create a folder to hold the experimental results
mkdir -p results. - Run
python ./perfect_match/apps/run_all_experiments.py ./perfect_match/apps news_treatment_assignment ./data ./results- The script will print all the command line configurations (1750 in total) you need to run to obtain the experimental results to reproduce the News results.
- Run the command line configurations from the previous step in a compute environment of your choice.
- After the experiments have concluded, use
./run_results.shto calculate the summary statistics mean +- standard deviation over all repeated runs in LaTeX syntax.- Example 1:
./run_results.sh ./results/pm_news8a5k_pbm_mse_1 news-8, wherenews-8indicates that you want results for the News-8 dataset and5kindicates kappa=5, to get the results for "PM" on News-8 with treatment assignment bias factor kappa set to 5. - Example 2:
./run_results.sh ./results/pm_news8a7k_pbm_mse_1 news-8to get the results for "PM" on News-8 with treatment assignment bias factor kappa set to 7. - Repeat for all evaluated methods / levels of kappa combinations.
- Example 1:
- Your results should match those found in the
perfect_match/visualisation/kappa_plot.Rfile.
TCGA Hidden Confounding (Figure 7)
- Navigate to the directory containing this file.
- Create a folder to hold the experimental results
mkdir -p results. - Run
python ./perfect_match/apps/run_all_experiments.py ./perfect_match/apps tcga ./data ./results- The script will print all the command line configurations (180 in total) you need to run to obtain the experimental results to reproduce the TCGA results.
- Run the command line configurations from the previous step in a compute environment of your choice.
- After the experiments have concluded, use
./run_results.shto calculate the summary statistics mean +- standard deviation over all repeated runs in LaTeX syntax.- Example 1:
./run_results.sh ./results/pm_tcga8a10k18478f_pbm_mse_1 tcga, wheretcgaindicates that you want results for the TCGA dataset and18478findicates the use of 18478 (out of 20531) features with the rest acting as hidden confounders, to get the results for "PM" with 10% hidden confounding on TCGA. - Example 2:
./run_results.sh ./results/pm_tcga8a10k16425f_pbm_mse_1 tcgato get the results for "PM" with 20% hidden confounding on TCGA. - Example 3:
./run_results.sh ./results/pm_tcga8a10k14372f_pbm_mse_1 tcgato get the results for "PM" with 30% hidden confounding on TCGA. - Repeat for all evaluated method / degree of hidden confounding combinations.
- Example 1:
- Your results should match those found in the
perfect_match/visualisation/confounding_plot.Rfile.
This work was partially funded by the Swiss National Science Foundation (SNSF) project No. 167302 within the National Research Program (NRP) 75 "Big Data". We gratefully acknowledge the support of NVIDIA Corporation with the donation of the Titan Xp GPUs used for this research. The results shown here are in whole or part based upon data generated by the TCGA Research Network: http://cancergenome.nih.gov/.